import matplotlib
if not hasattr(matplotlib.RcParams, "_get"):
matplotlib.RcParams._get = dict.get
Lecture 2: Variational Autoencoders (VAEs) from Scratch#
VAE 3D Visualization#
This notebook develops Variational Autoencoders (VAEs) from first principles.
We will cover:
The probabilistic prerequisites required for VAEs
Latent variable models
Why maximum likelihood becomes difficult
Variational inference and the approximate posterior
Full derivation of the ELBO
Expansion of the ELBO into reconstruction and KL terms
Why the reconstruction term becomes BCE or MSE depending on the decoder likelihood
The reparameterization trick
How encoder and decoder are trained jointly
Visual plots and code demonstrations for key concepts
1. High-Level Overview of VAEs#
A Variational Autoencoder (VAE) is a latent variable generative model.
It assumes that observed data \(x\) is generated using a hidden latent variable \(z\):
where:
\(z\) is a latent variable
\(p(z)\) is the prior over latents, usually a simple distribution such as \(\mathcal{N}(0, I)\)
\(p_\theta(x \mid z)\) is the decoder or generative model
\(\theta\) are the decoder parameters
The goals of a VAE are:
Learn a probabilistic generative model of the data
Learn meaningful latent representations
Enable generation of new samples by sampling \(z \sim p(z)\) and decoding it
The central difficulty is that learning requires the marginal likelihood
and this integral is generally intractable when \(p_\theta(x \mid z)\) is parameterized by a neural network.
That is why we need variational inference.
2. Probabilistic Prerequisites#
Before deriving VAEs, we need a few mathematical ingredients:
probability densities
Gaussian distributions
expectations
log-likelihood
KL divergence
Jensen’s inequality
latent variable models
posterior inference
2.1 Probability Densities#
For continuous random variables, probability is defined through a density.
If \(x\) has density \(p(x)\), then for any region \(A\),
For two random variables \(x\) and \(z\), the joint density is \(p(x, z)\), and we can write
The marginal density of \(x\) is
and the posterior is
2.2 Gaussian Distribution#
A univariate Gaussian is
A multivariate Gaussian in \(d\) dimensions is
A common VAE encoder uses a diagonal Gaussian:
This means the encoder outputs:
a mean vector \(\mu_\phi(x)\)
a variance vector \(\sigma_\phi^2(x)\)
def gaussian_pdf_1d(x, mu, sigma):
return (1.0 / (np.sqrt(2 * np.pi) * sigma)) * np.exp(-0.5 * ((x - mu) / sigma) ** 2)
x = np.linspace(-6, 6, 500)
plt.figure()
for mu, sigma in [(0, 1), (0, 0.5), (1.5, 1), (-1.5, 1.5)]:
plt.plot(x, gaussian_pdf_1d(x, mu, sigma), label=fr"$\mu={mu}, \sigma={sigma}$")
plt.title("1D Gaussian Distributions")
plt.xlabel("x")
plt.ylabel("Density")
plt.legend()
plt.grid(True, alpha=0.3)
plt.show()
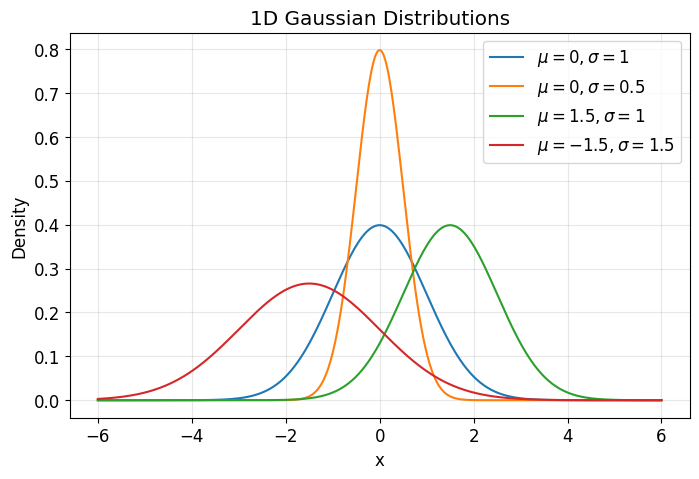
The plot above helps build intuition:
changing \(\mu\) shifts the distribution
changing \(\sigma\) changes spread or uncertainty
This becomes important in VAEs because the encoder maps each datapoint \(x\) to a Gaussian distribution in latent space.
2.3 Expectation#
For a random variable \(z \sim q(z)\) and a function \(f(z)\), the expectation is
For discrete variables,
Expectation is simply the average value of \(f(z)\) under the distribution \(q(z)\).
samples = rng.normal(loc=2.0, scale=1.5, size=100000)
empirical_mean = np.mean(samples)
empirical_second_moment = np.mean(samples**2)
print("Empirical E[z] =", empirical_mean)
print("Empirical E[z^2] =", empirical_second_moment)
print("Theoretical E[z] =", 2.0)
print("Theoretical E[z^2] =", 1.5**2 + 2.0**2)
Empirical E[z] = 1.993650604581616
Empirical E[z^2] = 6.241338452290414
Theoretical E[z] = 2.0
Theoretical E[z^2] = 6.25
The code above illustrates that expectation can be approximated using samples.
This is important because in VAEs the reconstruction term contains an expectation over \(q_\phi(z \mid x)\), which we often approximate using Monte Carlo sampling.
2.4 Log-Likelihood#
For a probabilistic model, we often want to maximize likelihood:
or equivalently maximize log-likelihood:
We prefer log-likelihood because:
it converts products into sums
it is numerically more stable
maximizing \(\log p_\theta(x)\) is equivalent to maximizing \(p_\theta(x)\) since \(\log\) is monotonic
For a dataset \(\{x^{(i)}\}_{i=1}^N\), maximum likelihood estimation solves
2.5 KL Divergence#
The KL divergence from \(q(z)\) to \(p(z)\) is
Important properties:
\(D_{\mathrm{KL}}(q\|p) \ge 0\)
\(D_{\mathrm{KL}}(q\|p) = 0\) if and only if \(q = p\) almost everywhere
it is not symmetric
In VAEs, the KL term measures how far the encoder posterior \(q_\phi(z \mid x)\) is from the prior \(p(z)\).
For two Gaussians where
the KL divergence is
where \(k\) is the dimension of \(z\).
If \(\Sigma = \operatorname{diag}(\sigma_1^2, \dots, \sigma_k^2)\), then
This is the standard closed-form VAE KL term.
def kl_diag_gaussian_to_standard_normal(mu, logvar):
"""
mu: array-like, shape (..., d)
logvar: array-like, shape (..., d)
returns KL for each sample
"""
return 0.5 * np.sum(np.exp(logvar) + mu**2 - 1.0 - logvar, axis=-1)
mu_values = np.linspace(-3, 3, 300)
logvar_fixed = np.zeros((300, 1)) # variance = 1
mu_grid = mu_values.reshape(-1, 1)
kl_vals = kl_diag_gaussian_to_standard_normal(mu_grid, logvar_fixed)
plt.figure()
plt.plot(mu_values, kl_vals)
plt.title(r"KL$(\mathcal{N}(\mu,1)\|\mathcal{N}(0,1))$ as a function of $\mu$")
plt.xlabel(r"$\mu$")
plt.ylabel("KL divergence")
plt.grid(True, alpha=0.3)
plt.show()
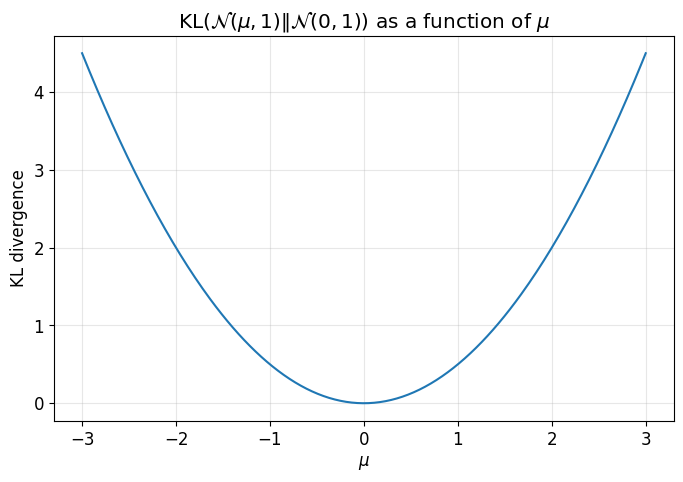
This plot shows that the KL divergence grows as the encoder mean moves away from the prior mean \(0\).
That is one way the KL term regularizes the latent space.
logvar_values = np.linspace(-4, 3, 300)
mu_fixed = np.zeros((300, 1))
logvar_grid = logvar_values.reshape(-1, 1)
kl_vals_var = kl_diag_gaussian_to_standard_normal(mu_fixed, logvar_grid)
plt.figure()
plt.plot(logvar_values, kl_vals_var)
plt.title(r"KL$(\mathcal{N}(0,\sigma^2)\|\mathcal{N}(0,1))$ as a function of $\log \sigma^2$")
plt.xlabel(r"$\log \sigma^2$")
plt.ylabel("KL divergence")
plt.grid(True, alpha=0.3)
plt.show()
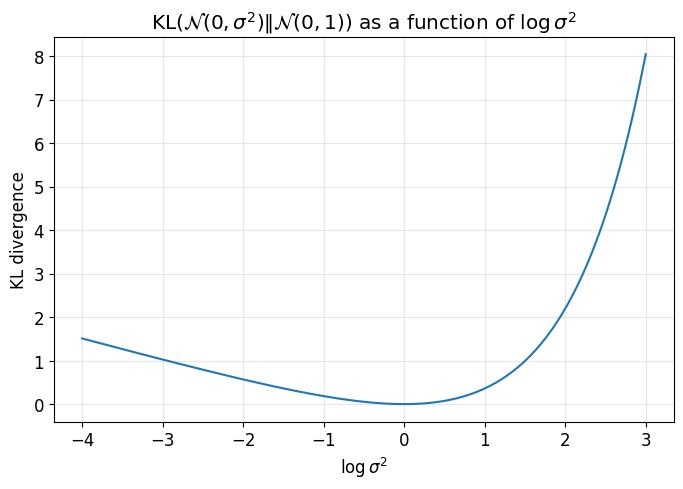
This second plot shows that the KL divergence is minimized when \(\sigma^2 = 1\), i.e. when \(\log \sigma^2 = 0\).
So the KL term encourages both:
mean near \(0\)
variance near \(1\)
2.6 Jensen’s Inequality#
For a concave function \(f\),
Since \(\log\) is concave,
This is the key inequality used to derive the ELBO as a lower bound.
# Simple illustration of Jensen's inequality for log
positive_samples = rng.uniform(0.1, 5.0, size=100000)
lhs = np.log(np.mean(positive_samples))
rhs = np.mean(np.log(positive_samples))
print("log(E[X]) =", lhs)
print("E[log X] =", rhs)
print("Difference log(E[X]) - E[log X] =", lhs - rhs)
log(E[X]) = 0.9341878920756458
E[log X] = 0.6874473352543539
Difference log(E[X]) - E[log X] = 0.24674055682129192
The output confirms:
This is exactly why the ELBO becomes a lower bound on \(\log p_\theta(x)\).
3. Latent Variable Models#
A latent variable model introduces hidden variables \(z\) to explain observed data \(x\).
The model is
and the marginal likelihood of a datapoint is
The latent variable \(z\) is intended to capture underlying structure that helps explain the data.
Why is exact maximum likelihood hard?#
We would ideally like to maximize
The problem is:
the integral over \(z\) is generally intractable
the posterior
is also intractable because it depends on the same marginal \(p_\theta(x)\)
So the model is elegant, but exact learning and inference are hard.
Variational Inference Idea#
To deal with the intractable posterior, we introduce an approximation:
where:
\(\phi\) are encoder parameters
\(q_\phi(z \mid x)\) is chosen to be tractable
we want it to approximate the true posterior \(p_\theta(z \mid x)\)
The central idea of VAEs is to optimize a tractable lower bound on \(\log p_\theta(x)\).
4. Autoencoder vs Variational Autoencoder#
A classical autoencoder does:
It learns a deterministic latent code \(h\).
A VAE instead learns a distribution over latent codes:
and then samples
before decoding through
So the VAE is probabilistic, and therefore generative.
5. ELBO Derivation from Scratch#
We begin with the log marginal likelihood:
We now multiply and divide by \(q_\phi(z \mid x)\) inside the integral:
Recognizing expectation under \(q_\phi(z \mid x)\), we get
Now apply Jensen’s inequality:
Therefore,
This lower bound is called the Evidence Lower Bound (ELBO):
So the ELBO is
and it satisfies
6. ELBO Derivation Using the True Posterior#
Now let us derive the same result in a way that makes the role of the posterior approximation clearer.
Consider the KL divergence between the approximate posterior and the true posterior:
Using Bayes’ rule,
So,
Since \(\log p_\theta(x)\) does not depend on \(z\),
Rearranging,
Therefore,
Since KL divergence is nonnegative,
and the bound becomes tight exactly when
7. Expanding the ELBO into the Standard VAE Form#
Recall:
Since the joint factorizes as
we have
Substituting this into the ELBO gives
Split the expectation:
The second term is exactly minus a KL divergence:
So the ELBO becomes
This is the standard VAE objective.
It has two terms:
Reconstruction term
\[ \mathbb{E}_{q_\phi(z \mid x)}[\log p_\theta(x \mid z)] \]which encourages good reconstructions
KL regularization term
\[ D_{\mathrm{KL}}(q_\phi(z \mid x)\|p(z)) \]which encourages the latent posterior to remain close to the prior
# Example parameters
prior_mean = np.array([0.0, 0.0])
prior_cov = np.array([
[1.0, 0.0],
[0.0, 1.0]
])
post_mean = np.array([1.5, -0.8])
post_cov = np.array([
[0.5, 0.0],
[0.0, 0.2]
])
# Plot
fig, ax = plt.subplots(figsize=(6, 6))
plot_gaussian_ellipse(
ax=ax,
mean=prior_mean,
cov=prior_cov,
n_std=2.0,
linestyle='-',
edgecolor='black',
linewidth=2,
)
plot_gaussian_ellipse(
ax=ax,
mean=post_mean,
cov=post_cov,
n_std=2.0,
linestyle='--',
edgecolor='black',
linewidth=2,
)
ax.scatter(*prior_mean, s=45)
ax.scatter(*post_mean, s=45)
ax.set_xlim(-4, 4)
ax.set_ylim(-4, 4)
ax.set_aspect("equal")
ax.set_title("Latent Regularization Intuition")
ax.set_xlabel(r"$z_1$")
ax.set_ylabel(r"$z_2$")
ax.grid(True, alpha=0.3)
ax.legend(handles=make_latent_regularization_legend(), loc='upper right')
plt.show()
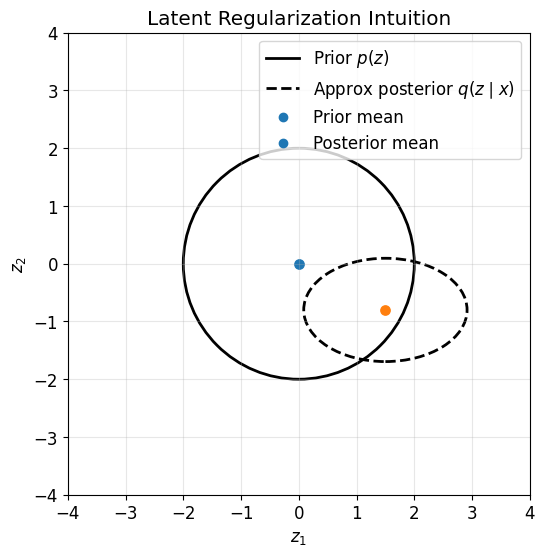
This figure illustrates the geometric role of the KL term.
The prior \(p(z)\) is centered at the origin with identity covariance
The approximate posterior \(q_\phi(z \mid x)\) for one datapoint may shift and shrink
The KL term penalizes large deviations from the prior
Thus the KL term organizes the latent space so that different datapoints live in a shared, regularized structure.
8. Why Maximizing the ELBO Makes Sense#
We ultimately want to maximize the true log-likelihood:
But from the identity
we see that maximizing the ELBO does two things:
it increases a lower bound on the true log-likelihood
it pushes the approximate posterior \(q_\phi(z \mid x)\) toward the true posterior \(p_\theta(z \mid x)\)
So ELBO maximization is a principled surrogate for maximum likelihood learning.
9. Encoder and Decoder Parameterization#
The encoder defines the approximate posterior:
So for each input \(x\), the encoder network outputs:
The decoder defines the conditional likelihood:
Its exact form depends on the type of data:
Bernoulli for binary data
Gaussian for continuous data
other likelihoods for other data types
So for each datapoint, the encoder maps \(x\) not to a single point, but to a distribution in latent space.
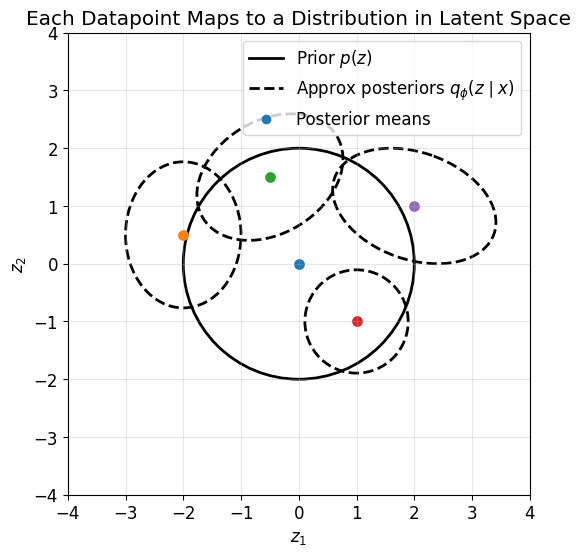
This plot gives an important conceptual picture:
each datapoint \(x\) is mapped by the encoder to a Gaussian
different datapoints correspond to different posterior distributions
the KL term prevents these from drifting arbitrarily far from the prior
10. The Reparameterization Trick#
The reconstruction term contains an expectation over samples from the encoder distribution:
We want gradients with respect to encoder parameters \(\phi\).
But naively sampling
creates a stochastic node that is not directly amenable to standard backpropagation.
To fix this, we use the reparameterization trick.
If
then we can sample by first drawing
and then setting
Now the randomness is isolated in \(\epsilon\), and \(z\) is a differentiable function of the encoder outputs.
mu = np.array([1.5, -0.5])
sigma = np.array([0.8, 0.3])
eps = rng.normal(size=(2000, 2))
z_samples = mu + sigma * eps
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
axes[0].scatter(eps[:, 0], eps[:, 1], s=8, alpha=0.3, label=r"$\epsilon \sim \mathcal{N}(0,I)$")
axes[0].set_title("Before Reparameterization")
axes[0].set_xlabel(r"$\epsilon_1$")
axes[0].set_ylabel(r"$\epsilon_2$")
axes[0].grid(True, alpha=0.3)
axes[0].legend()
axes[1].scatter(z_samples[:, 0], z_samples[:, 1], s=8, alpha=0.3, label=r"$z = \mu + \sigma \odot \epsilon$")
axes[1].scatter(*mu, s=80, label=r"$\mu$")
axes[1].set_title("After Reparameterization")
axes[1].set_xlabel(r"$z_1$")
axes[1].set_ylabel(r"$z_2$")
axes[1].grid(True, alpha=0.3)
axes[1].legend()
plt.tight_layout()
plt.show()
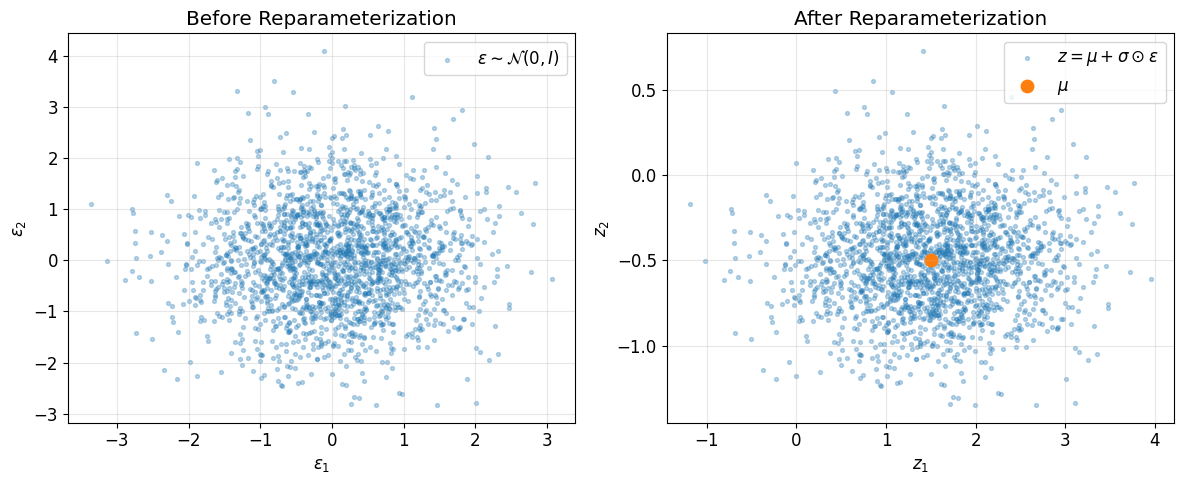
These two plots show the reparameterization geometrically:
first we sample standard Gaussian noise \(\epsilon\)
then we shift and scale it using the encoder outputs
this produces samples from the encoder posterior while keeping the computation differentiable
This is what allows gradients from the reconstruction term to flow back into the encoder.
11. Monte Carlo Approximation of the ELBO#
The expectation term
is usually approximated using Monte Carlo samples.
Using the reparameterization trick, we write
Then
In practice, during minibatch training, we usually take \(L=1\).
# Monte Carlo estimate of E[z^2] for z ~ N(mu, sigma^2)
mu = 1.0
sigma = 2.0
true_value = sigma**2 + mu**2
sample_sizes = [1, 5, 20, 100, 1000]
estimates = []
for n in sample_sizes:
eps = rng.normal(size=n)
z = mu + sigma * eps
estimates.append(np.mean(z**2))
print("True E[z^2] =", true_value)
for n, est in zip(sample_sizes, estimates):
print(f"MC estimate with {n:4d} samples = {est:.4f}")
True E[z^2] = 5.0
MC estimate with 1 samples = 1.2273
MC estimate with 5 samples = 8.6895
MC estimate with 20 samples = 9.0430
MC estimate with 100 samples = 4.6797
MC estimate with 1000 samples = 4.6531
This illustrates a general fact:
expectations can be estimated using samples
more samples usually reduce estimation noise
in VAEs, even a single sample often works well enough for SGD training
12. Closed-Form KL Term Used in VAEs#
For the common encoder choice
and prior
the KL divergence is
If the network outputs \(\log \sigma_j^2\), then since \(\sigma_j^2 = e^{\log \sigma_j^2}\), the KL becomes
This is the exact formula implemented in most VAE code.
13. Why the Reconstruction Term Becomes BCE or MSE#
The reconstruction term in the ELBO is
When we minimize negative ELBO, the reconstruction loss is
So the exact reconstruction loss depends entirely on the decoder likelihood.
13.1 Bernoulli Decoder#
If the data is binary or normalized to \([0,1]\), one common choice is
Then
Therefore,
So a Bernoulli decoder leads to binary cross-entropy reconstruction loss.
13.2 Gaussian Decoder with Fixed Variance#
If the data is continuous and we choose
with fixed \(\sigma_x^2\), then
Thus,
If \(\sigma_x^2\) is fixed, the constant does not affect optimization, so maximizing the likelihood is equivalent to minimizing mean squared error.
That is why MSE appears in many VAE implementations.
x_true = np.array([1.0, -0.5, 0.7])
xhat_candidates = np.array([
[1.0, -0.5, 0.7],
[0.8, -0.4, 0.9],
[0.0, 0.0, 0.0],
[2.0, -1.0, 1.4]
])
sigma2 = 0.25
def neg_log_gaussian_fixed_var(x, mu, sigma2):
D = len(x)
return 0.5 * D * np.log(2 * np.pi * sigma2) + (0.5 / sigma2) * np.sum((x - mu)**2)
print("Candidate reconstructions:")
for i, xhat in enumerate(xhat_candidates, 1):
mse = np.mean((x_true - xhat)**2)
nll = neg_log_gaussian_fixed_var(x_true, xhat, sigma2)
print(f"{i}: MSE={mse:.4f}, Gaussian NLL={nll:.4f}")
Candidate reconstructions:
1: MSE=0.0000, Gaussian NLL=0.6774
2: MSE=0.0300, Gaussian NLL=0.8574
3: MSE=0.5800, Gaussian NLL=4.1574
4: MSE=0.5800, Gaussian NLL=4.1574
The printed values show that when variance is fixed, lower MSE corresponds directly to higher Gaussian likelihood.
So MSE is not an arbitrary reconstruction loss. It comes from a probabilistic Gaussian decoder assumption.
13.3 Gaussian Decoder with Learned Variance#
If the decoder also predicts variance, then
and the log-likelihood becomes
This is no longer plain MSE.
Now the model can express uncertainty, and the loss includes:
a weighted squared error term
a log-variance penalty
13.4 Summary of Reconstruction Loss Choices#
In general,
So:
Bernoulli decoder \(\rightarrow\) BCE
Gaussian decoder with fixed variance \(\rightarrow\) MSE up to constants
Gaussian decoder with learned variance \(\rightarrow\) weighted MSE plus variance penalty
categorical decoder \(\rightarrow\) cross-entropy
The reconstruction loss is therefore dictated by the decoder likelihood assumption.
14. Final VAE Training Loss#
For one datapoint \(x\), the ELBO is
In practice we minimize the negative ELBO:
So the training loss is
For the standard Gaussian prior and diagonal Gaussian encoder,
where
15. How Encoder and Decoder Are Trained Together#
For one input \(x\), the computation proceeds as follows:
The encoder computes
\[ \mu_\phi(x), \qquad \log \sigma_\phi^2(x) \]Convert log variance to standard deviation:
\[ \sigma_\phi(x) = \exp\left(\frac{1}{2}\log \sigma_\phi^2(x)\right) \]Sample noise:
\[ \epsilon \sim \mathcal{N}(0,I) \]Reparameterize:
\[ z = \mu_\phi(x) + \sigma_\phi(x)\odot\epsilon \]Feed \(z\) into the decoder to get the parameters of \(p_\theta(x \mid z)\)
Compute the reconstruction term \(-\log p_\theta(x \mid z)\)
Compute the KL term
\[ D_{\mathrm{KL}}(q_\phi(z \mid x)\|p(z)) \]Add them to get the total loss
Backpropagate through the whole graph to update both encoder and decoder
The key point is:
the reconstruction term updates the decoder directly
through the reparameterized latent variable \(z\), the reconstruction term also updates the encoder
the KL term updates the encoder directly
So the encoder and decoder are trained jointly under a single objective.
Gradient Flow View#
The loss is
with
Then:
\(\nabla_\theta \mathcal{J}\) comes mainly from the decoder likelihood term
\(\nabla_\phi \mathcal{J}\) comes from:
the KL term directly
the reconstruction term indirectly through \(z\)
This indirect path works only because of the reparameterization trick.
16. Compact VAE Training Algorithm#
For each minibatch \(x\):
Compute encoder outputs
\[ \mu, \log \sigma^2 = \text{Encoder}_\phi(x) \]Sample noise
\[ \epsilon \sim \mathcal{N}(0,I) \]Reparameterize
\[ z = \mu + \exp\left(\frac{1}{2}\log \sigma^2\right)\odot \epsilon \]Decode
\[ \hat{x} \leftarrow \text{Decoder}_\theta(z) \]Compute reconstruction loss
\[ -\log p_\theta(x \mid z) \]Compute KL loss
\[ \frac{1}{2}\sum_j \left(\mu_j^2 + e^{\log\sigma_j^2} - \log\sigma_j^2 - 1\right) \]Total loss
\[ \mathcal{J} = \text{reconstruction loss} + \text{KL loss} \]Backpropagate and update \(\theta,\phi\)
# Toy single-datapoint VAE loss demo
# Fake datapoint
x = np.array([1.0, 0.0, 1.0, 1.0])
# Fake encoder outputs
mu = np.array([0.5, -0.3])
logvar = np.array([-0.2, 0.4])
# Sample latent
z, eps = sample_reparameterized(mu, logvar, rng)
# Fake decoder: simple linear map + sigmoid for Bernoulli parameters
W = np.array([
[1.2, -0.7, 0.5, 1.0],
[-0.4, 0.8, 1.1, -0.6]
])
b = np.array([0.1, -0.2, 0.0, 0.3])
logits = z @ W + b
xhat = sigmoid(logits)
recon_loss = binary_cross_entropy(x, xhat)
kl_loss = kl_diag_gaussian_to_standard_normal(mu.reshape(1, -1), logvar.reshape(1, -1))[0]
total_loss = recon_loss + kl_loss
print("x =", x)
print("mu =", mu)
print("logvar =", logvar)
print("sampled z =", z)
print("decoder xhat =", xhat)
print("recon loss =", recon_loss)
print("KL loss =", kl_loss)
print("total VAE loss =", total_loss)
x = [1. 0. 1. 1.]
mu = [ 0.5 -0.3]
logvar = [-0.2 0.4]
sampled z = [-0.80355319 0.88486174]
decoder xhat = [0.22825187 0.74466853 0.63912567 0.26221837]
recon loss = 4.628730077037161
KL loss = 0.22527772535962606
total VAE loss = 4.8540078023967865
This toy example numerically shows the two ingredients of the VAE loss:
reconstruction loss from the decoder likelihood
KL regularization from the encoder posterior
In a real neural implementation, the encoder and decoder would be neural networks, but the mathematical structure is exactly the same.
# A simple toy 1D decoder function to illustrate smooth latent structure
z_vals = np.linspace(-3, 3, 400)
decoded_mean = np.sin(1.5 * z_vals) + 0.2 * z_vals
plt.figure()
plt.plot(z_vals, decoded_mean)
plt.title("Toy Decoder Mean as a Function of Latent Variable z")
plt.xlabel("z")
plt.ylabel(r"Decoder output mean $\mu_\theta(z)$")
plt.grid(True, alpha=0.3)
plt.show()
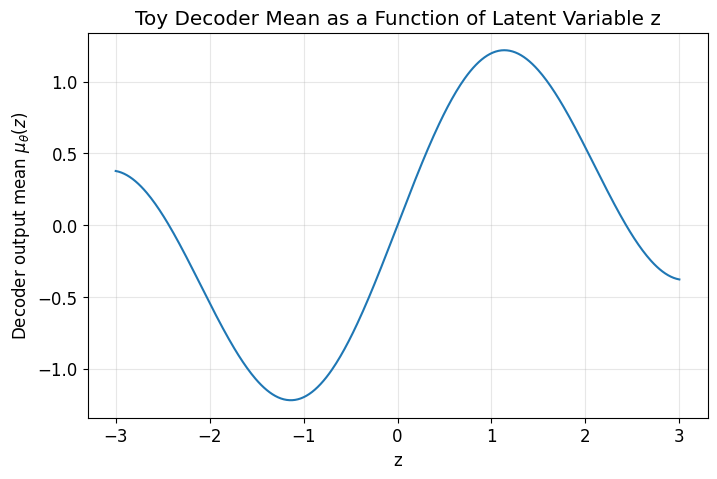
A VAE encourages the latent space to be smooth and structured.
The KL term helps ensure that latent points live in a regular region near the prior, and the decoder learns a smooth mapping from latent space to data space.
This is why nearby latent points often decode to similar outputs.
17. Why VAEs Are Generative Models#
After training, we can generate new samples without needing an input \(x\).
We simply:
sample from the prior
\[ z \sim p(z) = \mathcal{N}(0, I) \]decode
\[ x \sim p_\theta(x \mid z) \]
or use the decoder mean as the generated sample.
This is possible because the KL term aligned the encoder posteriors with the prior during training.
18. Important Practical and Conceptual Notes#
18.1 Why VAEs can produce blurry outputs#
A common reason is the decoder likelihood.
For example, a Gaussian decoder with independent pixels often encourages averaging across multiple plausible outputs, which can lead to blur.
18.2 Posterior collapse#
Sometimes the encoder posterior becomes too close to the prior:
for all \(x\).
Then the latent variable is effectively ignored by the decoder.
This often happens when the decoder is very powerful.
18.3 Beta-VAE#
A common variant is
or equivalently the loss
\(\beta > 1\) gives stronger regularization
\(\beta < 1\) gives weaker regularization
19. Final Summary#
A Variational Autoencoder is a latent variable model with:
a prior \(p(z)\)
an approximate posterior or encoder \(q_\phi(z \mid x)\)
a decoder likelihood \(p_\theta(x \mid z)\)
The true log-likelihood \(\log p_\theta(x)\) is generally intractable, so we optimize the ELBO:
Training minimizes the negative ELBO:
The reconstruction term encourages accurate decoding, while the KL term shapes the latent space so that sampling from the prior becomes meaningful.
The reparameterization trick
makes the whole model trainable by backpropagation.
That is the mathematical core of VAEs.
From scratch implementation of VAE (Live Code)#
# ----------------------------
# Run everything
# ----------------------------
X, y = make_pinwheel(points_per_class=800, num_classes=5, seed=10)
params = init_vae(input_dim=2, hidden_dim=64, latent_dim=2, seed=1)
# Train with visible progress
epochs = 800
batch_size = 256
lr = 3e-3
beta = 0.25
sigma2 = 0.03
seed = 11
rng_train = np.random.default_rng(seed)
m, v = init_adam(params)
n = len(X)
hist = {"loss": [], "recon": [], "kl": []}
t = 0
print("Starting VAE training...")
print(f"Dataset size: {n}")
print(f"Epochs: {epochs}, Batch size: {batch_size}, Learning rate: {lr}, beta: {beta}, sigma2: {sigma2}")
print("-" * 90)
for epoch in range(epochs):
idx = rng_train.permutation(n)
Xs = X[idx]
loss_sum = recon_sum = kl_sum = 0.0
batches = 0
for i in range(0, n, batch_size):
xb = Xs[i:i+batch_size]
t += 1
loss, recon, kl, grads = vae_step(xb, params, rng_train, beta=beta, sigma2=sigma2)
adam_step(params, grads, m, v, t, lr=lr)
loss_sum += loss
recon_sum += recon
kl_sum += kl
batches += 1
epoch_loss = loss_sum / batches
epoch_recon = recon_sum / batches
epoch_kl = kl_sum / batches
hist["loss"].append(epoch_loss)
hist["recon"].append(epoch_recon)
hist["kl"].append(epoch_kl)
if epoch == 0 or (epoch + 1) % 200 == 0 or epoch == epochs - 1:
print(
f"Epoch {epoch + 1:4d}/{epochs} | "
f"Total Loss: {epoch_loss:10.4f} | "
f"Recon: {epoch_recon:10.4f} | "
f"KL: {epoch_kl:8.4f}"
)
print("-" * 90)
print("Training complete.")
# Encode dataset into latent means
mu, logvar = encode_mean(X, params)
# Sample from prior until we get a decoded point that lies close to the learned data cloud
rng = np.random.default_rng(99)
z_new, x_new = None, None
best_z, best_x, best_d = None, None, 1e9
for _ in range(300):
z_try = rng.normal(size=(1, 2))
x_try = decode_mean(z_try, params)
d = nearest_dist_to_cloud(x_try[0], X)
if d < best_d:
best_d, best_z, best_x = d, z_try, x_try
if d < 0.18:
z_new, x_new = z_try, x_try
break
if z_new is None:
z_new, x_new = best_z, best_x
# A few additional generated samples for context
Z_gen = rng.normal(size=(400, 2))
X_gen = decode_mean(Z_gen, params)
# ----------------------------
# Plot
# ----------------------------
fig, axes = plt.subplots(2, 2, figsize=(13, 11))
# 1) input distribution
axes[0, 0].scatter(X[:, 0], X[:, 1], c=y, s=7, alpha=0.65, cmap="tab10")
axes[0, 0].set_title("Input 2D Data Distribution")
axes[0, 0].set_xlabel(r"$x_1$")
axes[0, 0].set_ylabel(r"$x_2$")
axes[0, 0].grid(True, alpha=0.25)
axes[0, 0].set_aspect("equal")
# 2) latent feature map
axes[0, 1].scatter(mu[:, 0], mu[:, 1], c=y, s=7, alpha=0.65, cmap="tab10")
axes[0, 1].scatter(z_new[0, 0], z_new[0, 1], s=180, marker="*", edgecolor="black", linewidth=1.2)
axes[0, 1].set_title("Latent Feature Map (Encoder Means)")
axes[0, 1].set_xlabel(r"$z_1$")
axes[0, 1].set_ylabel(r"$z_2$")
axes[0, 1].grid(True, alpha=0.25)
axes[0, 1].set_aspect("equal")
# 3) decoded sample overlaid on data
axes[1, 0].scatter(X[:, 0], X[:, 1], c="lightgray", s=7, alpha=0.45, label="training data")
axes[1, 0].scatter(X_gen[:, 0], X_gen[:, 1], s=8, alpha=0.18, label="decoded prior samples")
axes[1, 0].scatter(
x_new[0, 0], x_new[0, 1],
s=180, marker="*", edgecolor="black", linewidth=1.2,
label="decoded sampled point"
)
axes[1, 0].set_title("Sampled Latent Point Decoded Back to Data Space")
axes[1, 0].set_xlabel(r"$x_1$")
axes[1, 0].set_ylabel(r"$x_2$")
axes[1, 0].grid(True, alpha=0.25)
axes[1, 0].legend(loc="best")
axes[1, 0].set_aspect("equal")
# 4) training curves
axes[1, 1].plot(hist["loss"], label="total loss")
axes[1, 1].plot(hist["recon"], label="recon")
axes[1, 1].plot(hist["kl"], label="kl")
axes[1, 1].set_title("Training Curves")
axes[1, 1].set_xlabel("Epoch")
axes[1, 1].grid(True, alpha=0.25)
axes[1, 1].legend()
plt.tight_layout()
plt.show()
print("\nFinal sampled result:")
print("Sampled latent point z* =", np.round(z_new[0], 4))
print("Decoded data point x* =", np.round(x_new[0], 4))
print("Distance to nearest training point =", round(nearest_dist_to_cloud(x_new[0], X), 4))
Starting VAE training...
Dataset size: 4000
Epochs: 800, Batch size: 256, Learning rate: 0.003, beta: 0.25, sigma2: 0.03
------------------------------------------------------------------------------------------
Epoch 1/800 | Total Loss: 50.7059 | Recon: 49.8169 | KL: 3.5561
Epoch 200/800 | Total Loss: 1.8016 | Recon: 0.3068 | KL: 5.9793
Epoch 400/800 | Total Loss: 1.7483 | Recon: 0.2692 | KL: 5.9165
Epoch 600/800 | Total Loss: 1.7470 | Recon: 0.3017 | KL: 5.7812
Epoch 800/800 | Total Loss: 1.6717 | Recon: 0.3027 | KL: 5.4758
------------------------------------------------------------------------------------------
Training complete.
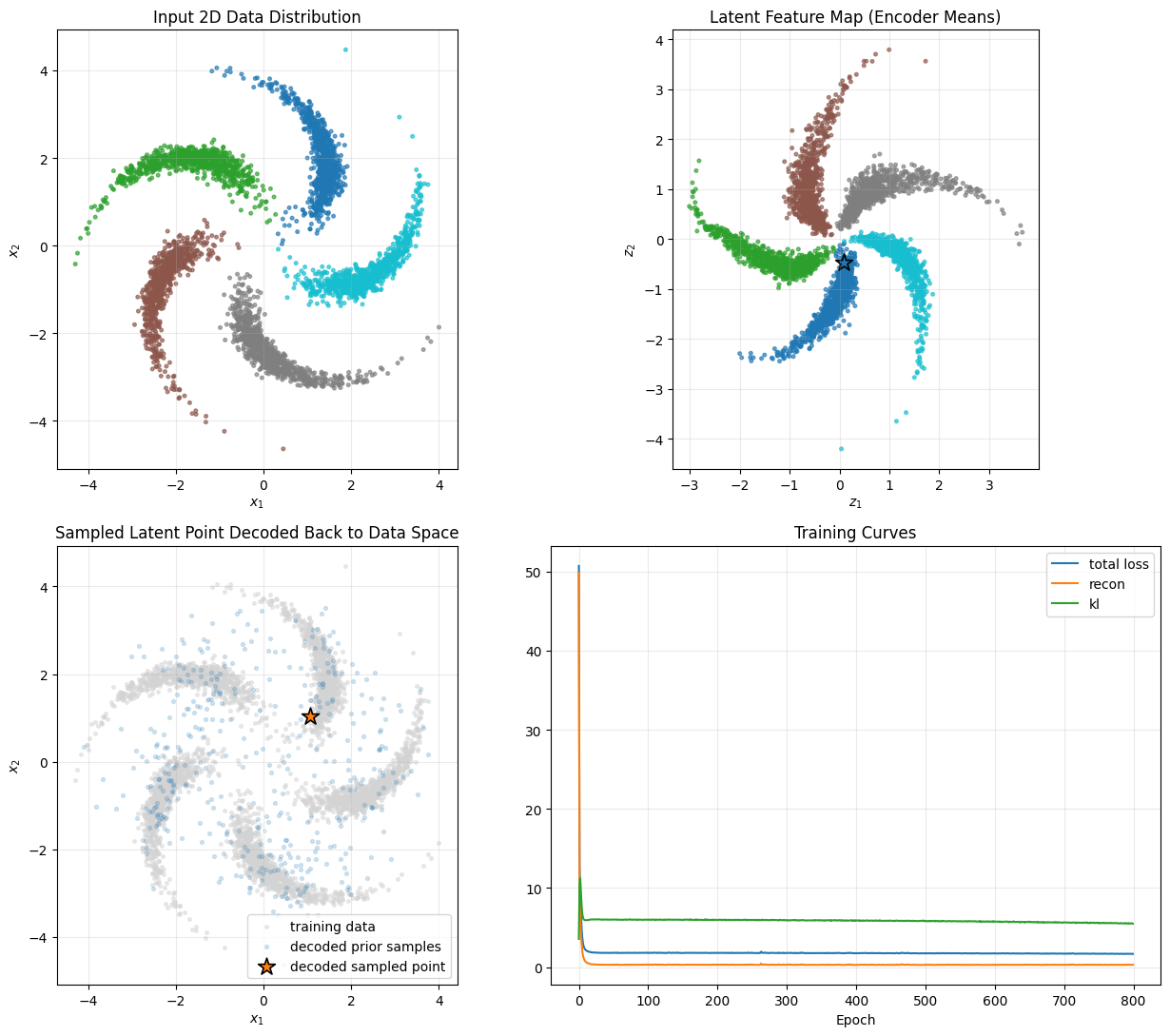
Final sampled result:
Sampled latent point z* = [ 0.0825 -0.4644]
Decoded data point x* = [1.0579 1.0317]
Distance to nearest training point = 0.0391

