import matplotlib
if not hasattr(matplotlib.RcParams, "_get"):
matplotlib.RcParams._get = dict.get
Lecture 3: Denoising Diffusion Probabilistic Models (DDPMs)#
This notebook is a mathematically rigorous, structured tutorial on Denoising Diffusion Probabilistic Models (DDPMs).
Goals of this notebook:
Build all prerequisites needed for DDPMs.
Derive the DDPM forward process and reverse process carefully.
Derive the variational objective and the practical training objective.
Connect the theory to implementation and visualization.
Provide plots and toy demonstrations for intuition.
We will use the following notation throughout:
\(x_0\): clean data sample
\(x_t\): latent/noisy sample at diffusion step \(t\)
\(q(\cdot)\): fixed forward/noising distributions
\(p_\theta(\cdot)\): learned reverse/generative distributions
\(\beta_t\): forward variance schedule
\(\alpha_t := 1 - \beta_t\)
\(\bar{\alpha}_t := \prod_{s=1}^t \alpha_s\), with \(\bar{\alpha}_0 = 1\)
We assume data lives in \(\mathbb{R}^d\).
Notebook roadmap#
This notebook is organized as follows:
Prerequisites
Diffusion intuition
Forward process derivation
Closed form for \(q(x_t \mid x_0)\)
Posterior \(q(x_{t-1} \mid x_t, x_0)\)
Reverse process model
ELBO derivation
Practical DDPM loss
\(\epsilon\)-prediction, \(x_0\)-prediction, and \(v\)-prediction
Training algorithm
Sampling algorithm
Implementation notes
Common confusions
Summary
Throughout, markdown cells contain the theory and derivations, while code cells are used for helper utilities and visual demonstrations.
Prerequisites#
Before deriving DDPMs, we need a strong grasp of:
probability densities and conditional distributions
Gaussian and multivariate Gaussian distributions
expectation and variance
KL divergence
change of variables
Markov chains
latent variable models
variational inference and ELBO
score function intuition
Gaussian identities
reparameterization-style identities
We now go through these carefully.
1. Probability densities and conditional distributions#
Let \(x \in \mathbb{R}^d\) be a continuous random variable with density \(p(x)\). Then
For two random variables \(x\) and \(y\), the joint density is \(p(x,y)\), and the conditional density is
whenever \(p(y) > 0\), where
The product rule gives
For a sequence \(x_{0:T} := (x_0, x_1, \dots, x_T)\),
A Markov chain is the special case where
Then
This is exactly the structure used in diffusion models.
2. Gaussian distributions#
2.1 Univariate Gaussian#
A scalar Gaussian with mean \(\mu\) and variance \(\sigma^2\) is
2.2 Multivariate Gaussian#
In \(\mathbb{R}^d\), a Gaussian with mean \(\mu \in \mathbb{R}^d\) and covariance \(\Sigma \in \mathbb{R}^{d \times d}\) is
In DDPMs, we frequently use isotropic covariances such as \(\sigma^2 I\).
3. Expectation and variance#
For a density \(p(x)\), the expectation of a function \(f(x)\) is
The mean is
The covariance is
For a scalar variable, variance is
Useful identity:
Linearity of expectation:
These become important when we manipulate Gaussian noise processes in diffusion.
4. KL divergence#
For two densities \(q(x)\) and \(p(x)\), the KL divergence is
Important facts:
\(D_{\mathrm{KL}}(q \| p) \ge 0\)
\(D_{\mathrm{KL}}(q \| p) = 0\) iff \(q=p\) almost everywhere
it is not symmetric
For Gaussians,
the KL divergence is
This is crucial in DDPMs because the ELBO decomposes into KL terms between Gaussian distributions.
5. Change of variables#
If \(z = f(x)\) is invertible and differentiable, then
Equivalently,
Diffusion models are not usually presented as normalizing flows, but the mindset that densities transform under mappings is still useful, especially when thinking about how noisy variables relate to clean variables.
6. Markov chains#
A sequence \(x_0, x_1, \dots, x_T\) is a Markov chain if
Then
Diffusion models use two Markov chains:
a forward chain \(q(x_{1:T} \mid x_0)\) that gradually adds noise
a reverse chain \(p_\theta(x_{0:T})\) that gradually removes noise
7. Latent variable models#
A latent variable model introduces hidden variables \(z\):
The marginal likelihood is
This integral is often intractable.
Diffusion models can be viewed as a latent variable model with many latent variables:
Here, \(x_0\) is observed data and the intermediate noisy states are latent variables.
8. Variational inference and ELBO#
Suppose we want to maximize \(\log p_\theta(x)\), but marginalization over latent variables is hard. Introduce an approximate posterior \(q(z \mid x)\):
Applying Jensen’s inequality,
This lower bound is the ELBO:
Also,
In DDPMs, the latent variables are \(x_{1:T}\) and the forward diffusion process \(q(x_{1:T}\mid x_0)\) plays the role of the variational distribution.
9. Score function intuition#
The score of a density \(p(x)\) is
It tells us the direction in which the density increases most rapidly.
In diffusion, denoising is closely related to learning the score of noisy data distributions. Later we will see that predicting the added noise \(\epsilon\) is closely tied to predicting the score.
10. Why Gaussian identities matter#
DDPM derivations rely on the following Gaussian facts:
Linear combinations of Gaussians are Gaussian.
Adding Gaussian noise to a Gaussian yields another Gaussian.
Products of Gaussian densities are proportional to Gaussians.
Conditionals of jointly Gaussian variables are Gaussian.
These facts are what make the following quantities analytically tractable:
\(q(x_t \mid x_0)\)
\(q(x_{t-1} \mid x_t, x_0)\)
the KL terms in the ELBO
11. Reparameterization-style identities#
If \(\epsilon \sim \mathcal{N}(0,I)\), then
has distribution
More generally, if \(LL^\top = \Sigma\), then
implies
This is the style of identity we repeatedly use in diffusion when writing noisy variables as clean signal plus scaled Gaussian noise.
Diffusion intuition#
We now step back and understand the big picture before doing the full derivation.
What problem diffusion models solve#
We want to model a complicated data distribution \(p_{\text{data}}(x)\) and generate new samples from it.
Direct generation in one shot is hard because the data distribution is high-dimensional and highly structured.
Diffusion models solve this by breaking the problem into many small denoising steps:
Define a simple process that gradually corrupts data into Gaussian noise.
Learn to reverse that process one small step at a time.
This turns a hard generation problem into a sequence of easier local denoising problems.
How diffusion differs from VAEs and GANs#
VAEs#
VAEs are latent variable models trained via an ELBO. They are principled and likelihood-based, but often produce blurrier samples.
GANs#
GANs can generate sharp samples but training can be unstable and they do not directly optimize likelihood.
DDPMs#
DDPMs are likelihood-based latent-variable generative models trained using a denoising objective closely tied to a variational bound. They are stable to train and generate high-quality samples by iterative refinement.
Forward diffusion vs reverse diffusion#
The forward diffusion process gradually adds noise:
By the end, \(x_T\) is approximately standard Gaussian noise.
The reverse diffusion process learns to invert this corruption:
At generation time, we start from Gaussian noise and repeatedly denoise.
The next plot shows the role of the noise schedule. This is one of the most important practical ingredients in diffusion.
We will visualize:
\(\beta_t\): the noise added at each step
\(\alpha_t = 1 - \beta_t\): the signal retention at each step
\(\bar{\alpha}_t = \prod_{s=1}^t \alpha_s\): the cumulative signal retention
T = 200
betas = linear_beta_schedule(T)
q = compute_diffusion_quantities(betas)
fig, ax = plt.subplots(figsize=(9, 5))
ax.plot(q["betas"], label=r'$\beta_t$')
ax.plot(q["alphas"], label=r'$\alpha_t = 1-\beta_t$')
ax.plot(q["alpha_bars"], label=r'$\bar{\alpha}_t$')
ax.set_title("Diffusion schedule quantities")
ax.set_xlabel("timestep t")
ax.set_ylabel("value")
ax.legend()
ax.grid(True, alpha=0.3)
plt.show()
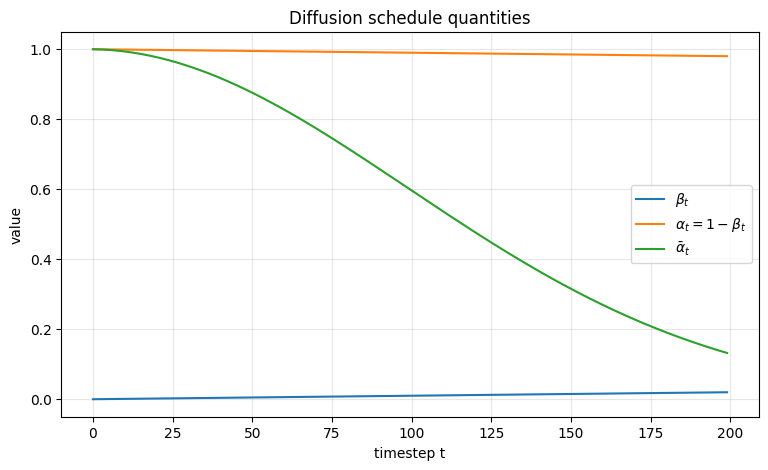
Interpretation of the plot#
\(\beta_t\) determines how much fresh noise is injected at each step.
\(\alpha_t\) determines how much of the previous signal survives each step.
\(\bar{\alpha}_t\) tells us how much of the original signal \(x_0\) survives after \(t\) steps.
Later we will show that
so \(\bar{\alpha}_t\) is the central cumulative quantity in DDPMs.
Forward process derivation#
We now derive the forward diffusion process carefully.
Definition of the forward process#
Choose a variance schedule \(\{\beta_t\}_{t=1}^T\) with \(\beta_t \in (0,1)\) and define
The forward process is defined by
Equivalently,
So each step:
shrinks the previous signal by \(\sqrt{\alpha_t}\)
adds fresh Gaussian noise of variance \(\beta_t I\)
Why this parameterization is useful#
The parameterization
is convenient because:
it keeps the process Gaussian
it separates signal retention and noise injection
it gives clean cumulative formulas in terms of \(\bar{\alpha}_t\)
The full forward chain factorizes as
The next demonstration visualizes the forward noising process on a simple 1D signal. This is not meant to be a realistic image diffusion example; it is purely pedagogical so we can see how the signal is gradually destroyed by noise.
# Better visualization of forward noising:
# 1) separate subplots for selected timesteps
# 2) heatmap of the whole trajectory over time
# 3) signal/noise coefficients across timesteps
import numpy as np
import matplotlib.pyplot as plt
# --- simple clean 1D signal ---
n = 300
x_axis = np.linspace(0, 2 * np.pi, n)
x0_signal = np.sin(x_axis) + 0.3 * np.sin(4 * x_axis)
# --- diffusion schedule ---
T_demo = 100
betas_demo = linear_beta_schedule(T_demo, 1e-4, 2e-2)
sched_demo = compute_diffusion_quantities(betas_demo)
# --- generate one forward trajectory iteratively ---
rng = np.random.default_rng(42)
xs_demo = sample_forward_iterative(x0_signal, betas_demo, rng=rng) # shape: [T+1, n]
# ---------------------------------------------------
# Figure 1: separate subplots for selected timesteps
# ---------------------------------------------------
timesteps_to_show = [0, 10, 30, 60, 100]
fig, axes = plt.subplots(len(timesteps_to_show), 1, figsize=(10, 10), sharex=True)
for ax, t in zip(axes, timesteps_to_show):
ax.plot(x_axis, x0_signal, linestyle='--', alpha=0.8, label=r'clean $x_0$')
ax.plot(x_axis, xs_demo[t], label=fr'$x_{{{t}}}$')
ax.set_ylabel("value")
ax.set_title(f"Forward noising at timestep t={t}")
ax.grid(True, alpha=0.3)
ax.legend(loc="upper right")
axes[-1].set_xlabel("signal coordinate")
plt.tight_layout()
plt.show()
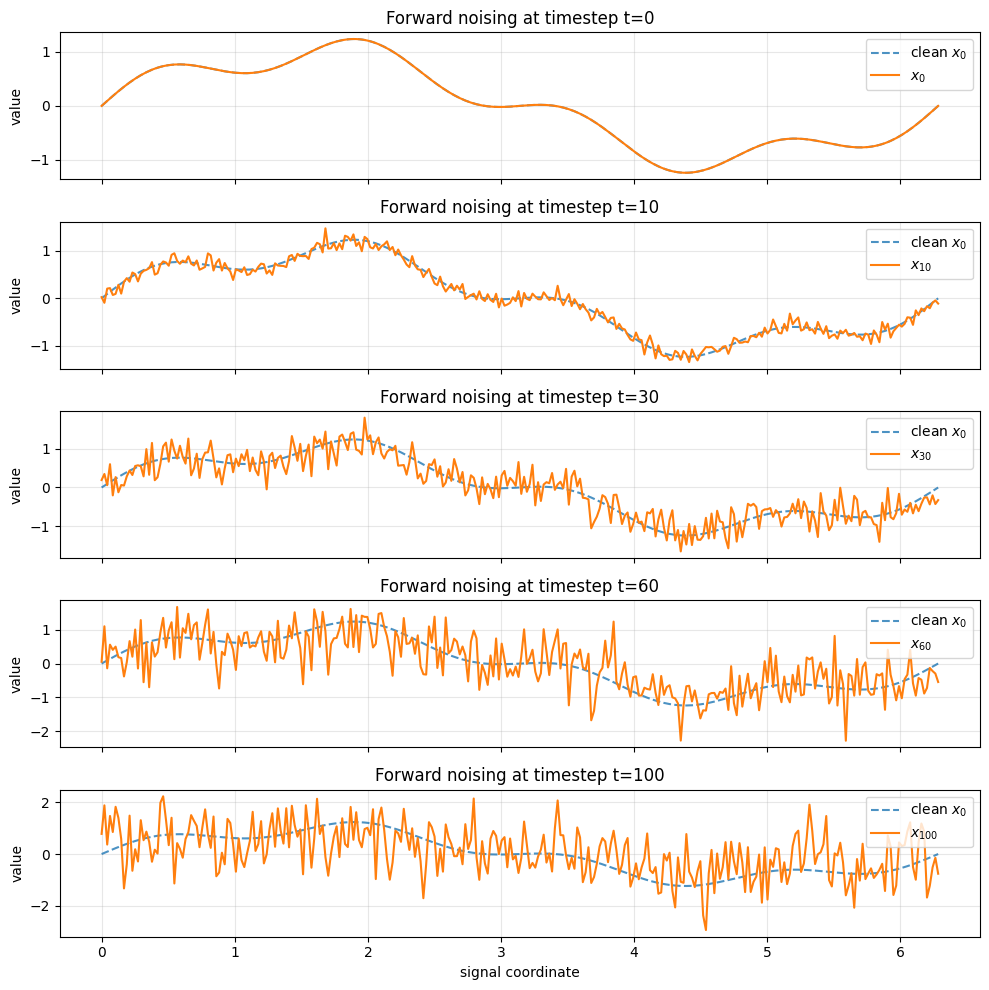
Interpretation#
At early timesteps, the signal still strongly resembles \(x_0\).
At later timesteps, signal structure is weakened and noise dominates.
This is exactly what the diffusion forward process is designed to do: transform complicated data into something close to standard Gaussian noise.
Closed form for \(q(x_t \mid x_0)\)#
This is one of the central derivations in DDPMs.
We will derive
where
Step-by-step derivation of \(q(x_t \mid x_0)\)#
From the one-step forward process,
Let us expand the first few cases.
For \(t=1\)#
So immediately,
Since \(\beta_1 = 1-\alpha_1\) and \(\bar{\alpha}_1 = \alpha_1\), this matches
For \(t=2\)#
Substitute the expression for \(x_1\):
Distribute:
Since this is a linear combination of independent Gaussians, it is Gaussian.
Its conditional mean is
Its conditional covariance is
Now simplify:
Hence
This suggests the general formula.
Inductive derivation#
Assume
Then we may write
Apply one forward step:
Substitute the representation for \(x_{t-1}\):
Distribute:
Since \(\bar{\alpha}_t = \alpha_t\bar{\alpha}_{t-1}\),
The last two terms are independent zero-mean Gaussians, so their sum is Gaussian with covariance
Simplify the scalar coefficient:
Since \(\alpha_t + \beta_t = 1\),
Therefore,
One-step sampling identity#
The previous result gives the very important identity
This means we can sample \(x_t\) directly from \(x_0\) in one step during training, without simulating all intermediate states.
This is one of the biggest practical simplifications in DDPM training.
Why \(\bar{\alpha}_t\) is so useful#
The quantity \(\bar{\alpha}_t\) captures the cumulative signal retention.
\(\sqrt{\bar{\alpha}_t}\) scales the clean signal \(x_0\)
\(\sqrt{1-\bar{\alpha}_t}\) scales the total accumulated noise
So \(\bar{\alpha}_t\) summarizes the full history of the forward process up to time \(t\).
The next code cell verifies numerically that iterative noising and the closed-form marginal have the same mean and variance structure.
T_check = 80
betas_check = linear_beta_schedule(T_check)
sched_check = compute_diffusion_quantities(betas_check)
x0_scalar = np.array([2.0])
target_t = 50
num_samples = 10000
rng = np.random.default_rng(123)
iterative_samples = []
closed_form_samples = []
for _ in range(num_samples):
x = x0_scalar.copy()
for s in range(target_t):
beta = betas_check[s]
alpha = 1.0 - beta
x = np.sqrt(alpha) * x + np.sqrt(beta) * rng.normal(size=x.shape)
iterative_samples.append(x.item())
for _ in range(num_samples):
eps = rng.normal(size=x0_scalar.shape)
x = np.sqrt(sched_check["alpha_bars"][target_t - 1]) * x0_scalar + np.sqrt(1.0 - sched_check["alpha_bars"][target_t - 1]) * eps
closed_form_samples.append(x.item())
iterative_samples = np.array(iterative_samples)
closed_form_samples = np.array(closed_form_samples)
theoretical_mean = np.sqrt(sched_check["alpha_bars"][target_t - 1]) * x0_scalar.item()
theoretical_var = 1.0 - sched_check["alpha_bars"][target_t - 1]
print("Iterative empirical mean:", iterative_samples.mean())
print("Closed-form empirical mean:", closed_form_samples.mean())
print("Theoretical mean:", theoretical_mean)
print()
print("Iterative empirical variance:", iterative_samples.var())
print("Closed-form empirical variance:", closed_form_samples.var())
print("Theoretical variance:", theoretical_var)
Iterative empirical mean: 1.7108073910406751
Closed-form empirical mean: 1.7115163810255865
Theoretical mean: 1.7086408359524738
Iterative empirical variance: 0.26582835854634257
Closed-form empirical variance: 0.2641044738782164
Theoretical variance: 0.2701366234289079
fig, ax = plt.subplots(figsize=(9, 5))
ax.hist(iterative_samples, bins=120, density=True, alpha=0.5, label='iterative')
ax.hist(closed_form_samples, bins=120, density=True, alpha=0.5, label='closed-form')
ax.set_title("Iterative noising vs closed-form noising at fixed t")
ax.set_xlabel("x_t")
ax.set_ylabel("density")
ax.legend()
ax.grid(True, alpha=0.3)
plt.show()
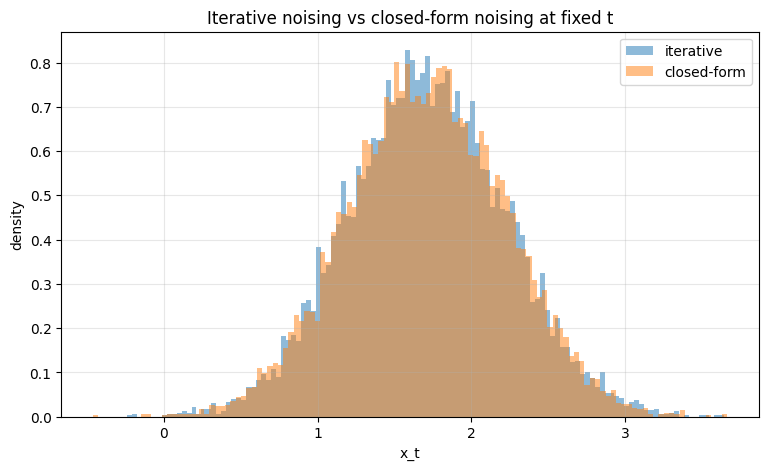
The histograms should overlap closely, confirming the derivation of
Posterior \(q(x_{t-1} \mid x_t, x_0)\)#
This posterior is the next crucial quantity in DDPMs.
We will derive that
with explicit formulas for \(\tilde{\mu}_t\) and \(\tilde{\beta}_t\).
Step 1: Bayes structure#
By the Markov property,
Since \(x_t\) depends only on \(x_{t-1}\),
So
Now plug in the two Gaussian factors:
As a function of \(x_{t-1}\), this is the product of two Gaussians, hence proportional to another Gaussian.
Step 2: Expand the exponent#
Ignoring constants independent of \(x_{t-1}\),
Expand the first quadratic:
Expand the second quadratic:
Collect only terms involving \(x_{t-1}\). The exponent becomes
This is the canonical form of a Gaussian.
Step 3: Read off the precision and mean#
Define
and
Then the Gaussian has variance
and mean
Step 4: Simplify the variance#
We have
Put over a common denominator:
Simplify the numerator:
Since \(\alpha_t + \beta_t = 1\),
Hence
Therefore,
Step 5: Simplify the mean#
We have
Substitute the simplified variance:
Distribute:
So the posterior is
where
and
The next cell visualizes how the posterior variance behaves across time. This variance is important because it tells us how uncertain the reverse transition remains when both \(x_t\) and \(x_0\) are known.
T_plot = 200
betas_plot = linear_beta_schedule(T_plot)
sched_plot = compute_diffusion_quantities(betas_plot)
fig, ax = plt.subplots(figsize=(9, 5))
ax.plot(sched_plot["betas"], label=r'$\beta_t$')
ax.plot(sched_plot["posterior_variance"], label=r'$\tilde{\beta}_t$ (posterior variance)')
ax.set_title("Forward variance vs posterior variance")
ax.set_xlabel("timestep t")
ax.set_ylabel("variance")
ax.legend()
ax.grid(True, alpha=0.3)
plt.show()
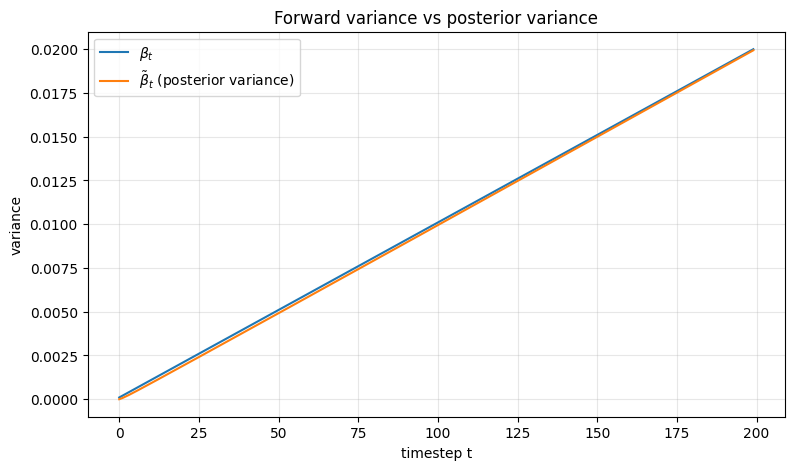
Interpretation#
\(\beta_t\) is the variance injected by the forward process
\(\tilde{\beta}_t\) is the variance of the posterior \(q(x_{t-1}\mid x_t, x_0)\)
The posterior variance is generally smaller because conditioning on both \(x_t\) and \(x_0\) gives more information about \(x_{t-1}\).
Reverse process model#
We now define the learned reverse generative process.
Learned reverse chain#
At generation time, we want to start from Gaussian noise and gradually denoise.
We define a simple prior at the terminal timestep:
Then define a learned reverse Markov chain
Each reverse conditional is parameterized as a Gaussian:
Typically we model the mean and either fix the variance or learn it.
Why Gaussian reverse transitions make sense#
Although \(q(x_{t-1}\mid x_t)\) is generally intractable, the conditional
has an exact Gaussian form. That suggests a Gaussian family for the learned reverse transition.
Also, because each forward step is small, the reverse transition is locally close to Gaussian. This is one of the key reasons DDPMs are tractable.
ELBO derivation#
We now derive the variational training objective for DDPMs.
Step 1: Marginal likelihood of the data#
For one observed data point \(x_0\), the model likelihood is
Introduce the forward process \(q(x_{1:T}\mid x_0)\):
This becomes
Apply Jensen’s inequality:
Define the ELBO:
Training minimizes the negative ELBO:
Step 2: Expand the factorization#
The forward process factorizes as
so
The reverse model factorizes as
so
Plugging into the ELBO gives
Step 3: Rewrite the forward chain in posterior form#
A key identity is
This comes from repeatedly applying Bayes’ rule to the forward chain.
Using that identity, we can rewrite the ratio inside the ELBO as
Taking logs and expectations yields the standard decomposition.
Step 4: ELBO decomposition across timesteps#
The negative ELBO can be written as
This is one of the most important equations in DDPM theory.
It has three parts:
A prior-matching term at the terminal time
A sum of timestep-wise KL terms
A reconstruction term at the first step
Interpretation of the ELBO terms#
Let
and
Then
\(L_T\) encourages the final noisy distribution to match a standard Gaussian
\(L_{t-1}\) trains the reverse model to match the true posterior transition
\(L_0\) is the final reconstruction-like term
In practice, the middle KL terms are the ones that lead to the familiar noise-prediction objective.
Practical DDPM loss#
We now derive the practical training objective used in DDPMs.
Step 1: Choose a Gaussian reverse model with fixed variance#
A common DDPM choice is
where \(\sigma_t^2\) is fixed.
The true posterior is
Since both are Gaussian, the KL term is analytically tractable. If the variances are fixed, then up to constants and known weights, minimizing the KL is equivalent to matching the means:
Step 2: Express the posterior mean using the added noise \(\epsilon\)#
Recall the closed-form noising identity
Solve this for \(x_0\):
Now substitute into
After algebraic simplification, one obtains
This is the key bridge from posterior mean matching to noise prediction.
Step 3: Parameterize the model via noise prediction#
Let the neural network predict the noise:
Then define the reverse mean as
Step 4: Show how the MSE on noise appears#
We have
and
Subtract:
Therefore,
Hence, each KL term is proportional to a weighted MSE between the true noise and the predicted noise.
Simplified DDPM objective#
The original DDPM paper found that a simplified unweighted loss works very well in practice:
where
This is the standard training objective used in many diffusion implementations.
The next code cell demonstrates the relationship between:
clean sample \(x_0\)
noisy sample \(x_t\)
noise \(\epsilon\)
and shows that if we know \(\epsilon\), we can recover \(x_0\) exactly through
rng = np.random.default_rng(7)
n = 400
x = np.linspace(0, 1, n)
x0_demo = np.sin(2 * np.pi * x) + 0.5 * np.cos(6 * np.pi * x)
T_demo2 = 100
betas_demo2 = linear_beta_schedule(T_demo2)
sched_demo2 = compute_diffusion_quantities(betas_demo2)
t_demo = 60
alpha_bar_t = sched_demo2["alpha_bars"][t_demo - 1]
eps = rng.normal(size=x0_demo.shape)
xt_demo = np.sqrt(alpha_bar_t) * x0_demo + np.sqrt(1 - alpha_bar_t) * eps
x0_recovered = recover_x0_from_epsilon_prediction(xt_demo, eps, alpha_bar_t)
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(x0_demo, label=r'true $x_0$')
ax.plot(xt_demo, label=r'noisy $x_t$', alpha=0.8)
ax.plot(x0_recovered, '--', label=r'recovered $x_0$ from true $\epsilon$')
ax.set_title("Recovering x0 from xt and epsilon")
ax.set_xlabel("signal coordinate")
ax.set_ylabel("value")
ax.legend()
ax.grid(True, alpha=0.3)
plt.show()
print("Max absolute reconstruction error:", np.max(np.abs(x0_demo - x0_recovered)))
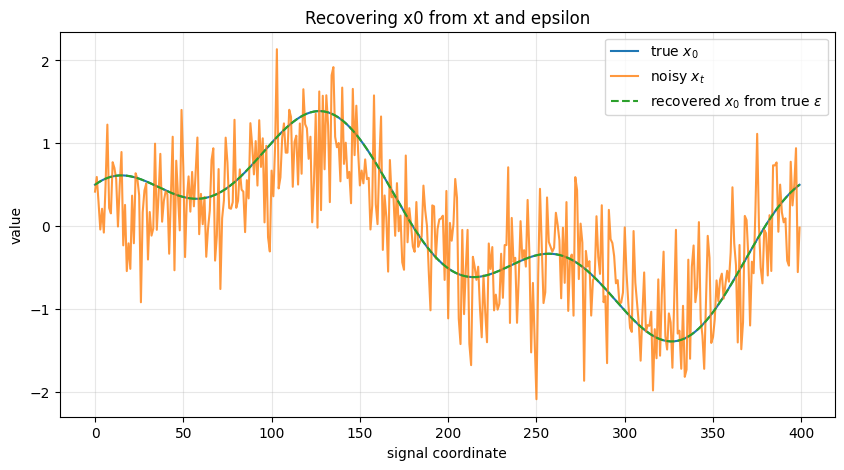
Max absolute reconstruction error: 2.220446049250313e-16
\(\epsilon\)-, \(x_0\)-, and \(v\)-parameterizations#
The reverse model can be parameterized in multiple equivalent ways.
1. \(\epsilon\)-prediction#
The network predicts the noise:
This is the standard DDPM parameterization.
Given \(\epsilon_\theta\), the reverse mean is
2. \(x_0\)-prediction#
Instead of predicting noise, the network may directly predict the clean sample:
Then the predicted noise can be reconstructed by
Or equivalently, one may plug \(\hat{x}_{0,\theta}\) directly into the posterior mean formula.
3. \(v\)-prediction#
A popular modern alternative is to predict
This is another linear reparameterization of the same information.
Given \((x_t, v)\), we can recover both \(x_0\) and \(\epsilon\):
So the three parameterizations are mathematically equivalent up to invertible linear transformations.
The next cell numerically verifies the equivalence between the parameterizations.
rng = np.random.default_rng(21)
x0 = rng.normal(size=1000)
eps = rng.normal(size=1000)
alpha_bar_t = 0.37
xt = np.sqrt(alpha_bar_t) * x0 + np.sqrt(1 - alpha_bar_t) * eps
v = v_from_x0_epsilon(x0, eps, alpha_bar_t)
x0_rec = x0_from_xt_v(xt, v, alpha_bar_t)
eps_rec = eps_from_xt_v(xt, v, alpha_bar_t)
print("max |x0 - x0_rec| =", np.max(np.abs(x0 - x0_rec)))
print("max |eps - eps_rec| =", np.max(np.abs(eps - eps_rec)))
max |x0 - x0_rec| = 4.440892098500626e-16
max |eps - eps_rec| = 4.440892098500626e-16
Training algorithm#
We now state the DDPM training procedure explicitly.
Training pipeline#
For each training iteration:
Sample a clean data point \(x_0 \sim p_{\text{data}}\).
Sample a timestep \(t\) uniformly from \(\{1,\dots,T\}\).
Sample noise \(\epsilon \sim \mathcal{N}(0,I)\).
Construct the noisy sample
Feed \((x_t, t)\) into the neural network.
Predict one of:
\(\epsilon\)
\(x_0\)
\(v\)
Compute the corresponding loss, most commonly
Update the network parameters by gradient descent.
Notice that training never needs to simulate the full forward chain step by step; direct noising via the closed form is enough.
Sampling algorithm#
We now state the generation procedure at inference time.
Reverse ancestral sampling#
At generation time we do not know \(x_0\).
Instead:
Sample
For \(t = T, T-1, \dots, 1\):
evaluate the neural network on \((x_t,t)\)
construct the reverse mean \(\mu_\theta(x_t,t)\)
sample
Return \(x_0\)
This is called ancestral sampling because we sample sequentially from the learned reverse Markov chain.
The next demo does not use a trained neural network. Instead, it uses the true \(\epsilon\) from the synthetic forward construction, so that we can see how the reverse mean behaves in an idealized setting.
rng = np.random.default_rng(1234)
n = 300
grid = np.linspace(0, 2 * np.pi, n)
x0_clean = np.sin(grid) + 0.3 * np.sin(5 * grid)
T_toy = 80
betas_toy = linear_beta_schedule(T_toy)
sched_toy = compute_diffusion_quantities(betas_toy)
t = 50
alpha_bar_t = sched_toy["alpha_bars"][t - 1]
eps_true = rng.normal(size=x0_clean.shape)
xt = np.sqrt(alpha_bar_t) * x0_clean + np.sqrt(1 - alpha_bar_t) * eps_true
mu_reverse = ddpm_reverse_mean_from_eps(
xt, eps_true, t, sched_toy["betas"], sched_toy["alphas"], sched_toy["alpha_bars"]
)
true_posterior_mean, true_posterior_var = posterior_mean_variance(
x0_clean, xt, t, sched_toy["betas"], sched_toy["alphas"], sched_toy["alpha_bars"]
)
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(xt, label=r'$x_t$', alpha=0.7)
ax.plot(mu_reverse, label='reverse mean from true epsilon')
ax.plot(true_posterior_mean, '--', label='closed-form posterior mean')
ax.set_title("Reverse mean computed two equivalent ways")
ax.set_xlabel("signal coordinate")
ax.set_ylabel("value")
ax.legend()
ax.grid(True, alpha=0.3)
plt.show()
print("Max difference between two means:", np.max(np.abs(mu_reverse - true_posterior_mean)))
print("Posterior variance scalar:", true_posterior_var)
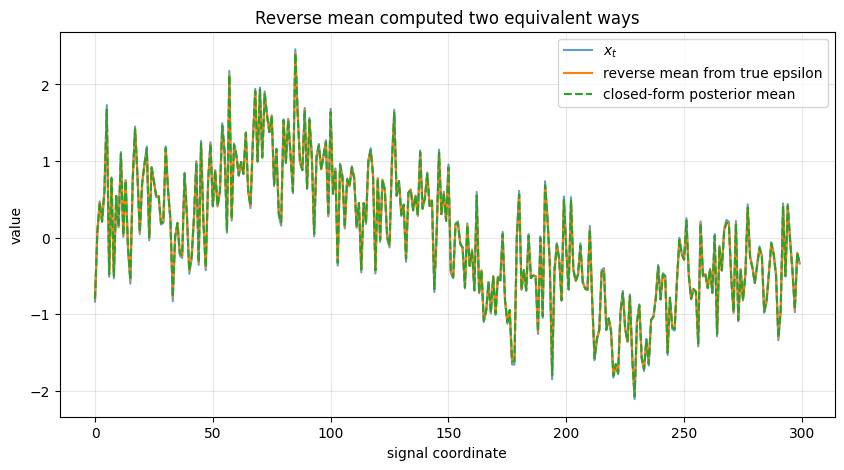
Max difference between two means: 4.440892098500626e-16
Posterior variance scalar: 0.012019444947095831
This verifies the equivalence between:
the posterior mean formula derived from Gaussian algebra
the mean formula expressed through \(\epsilon\)-prediction
That equivalence is the mathematical reason the DDPM training objective can be written as noise prediction.
Implementation notes#
We now connect the theory to actual implementation details.
What is actually sampled during training?#
For each minibatch item, we sample:
a clean data point \(x_0\)
a timestep \(t\)
Gaussian noise \(\epsilon\)
Then we construct
This pair \((x_t, t)\) becomes the network input, and \(\epsilon\) becomes the supervision target for standard DDPM training.
Why the network needs the timestep#
The amount of noise in \(x_t\) depends strongly on \(t\).
The same input pattern could correspond to very different denoising problems depending on whether the sample is only slightly noisy or almost pure noise.
So the network must know the timestep. In practice, \(t\) is embedded using sinusoidal or learned embeddings and injected into the network.
Why direct noising from \(x_0\) is enough during training#
The forward process is defined as a Markov chain, but the closed form
gives the exact marginal at time \(t\).
So training can skip explicit simulation of all intermediate forward states. This is mathematically exact, not an approximation.
Why the final prior is standard Gaussian#
If \(T\) is large enough and the schedule is chosen well, then repeated noising drives samples toward something close to
This justifies choosing
as the start of the reverse generative process.
Relationship to score matching#
For
the score with respect to \(x_t\) is
Since
we get
So predicting \(\epsilon\) is equivalent, up to a known scale, to predicting the score of the noisy conditional distribution.
Brief intuition for classifier-free guidance#
In conditional diffusion, the model may be trained both:
conditionally: \(\epsilon_\theta(x_t,t,c)\)
unconditionally: \(\epsilon_\theta(x_t,t,\varnothing)\)
Then at sampling time we use
where \(w\) is the guidance scale.
Interpretation:
the unconditional prediction gives the baseline denoising direction
the conditional prediction adds information specific to the condition
amplifying the difference strengthens adherence to the condition
Brief mention of DDIM#
DDPM uses stochastic ancestral sampling.
DDIM modifies the sampling process to allow deterministic or partially deterministic trajectories while using the same training objective.
So:
DDPM: stochastic reverse chain
DDIM: alternative non-Markovian sampling procedure derived from the same learned model
Common confusions and clarifications#
1. Why is \(q(x_{t-1}\mid x_t,x_0)\) tractable but \(q(x_{t-1}\mid x_t)\) intractable?#
Because conditioning on \(x_0\) makes the problem Gaussian and linear.
Without conditioning on \(x_0\), one would need to marginalize over the unknown data posterior:
and \(q(x_0\mid x_t)\) depends on the true data distribution, which is not analytically known.
2. Why predict \(\epsilon\) instead of directly predicting \(x_{t-1}\)?#
Because:
the target \(\epsilon\) has a simple standard Gaussian distribution
the posterior mean can be expressed directly in terms of \(\epsilon\)
the training objective becomes a simple regression problem
it exposes the connection to score matching
3. Does direct sampling of \(x_t\) break the Markov interpretation?#
No.
The forward chain still exists mathematically. The direct formula
is simply the exact closed-form marginal of that chain.
4. Is DDPM maximizing likelihood exactly?#
Not exactly. It optimizes an ELBO, which is a lower bound on the data log-likelihood.
Then the practical simplified objective is an especially convenient surrogate of that ELBO.
5. Where does denoising actually happen?#
At every reverse step.
The model sees a noisy sample \(x_t\), predicts the noise or the clean component, constructs a reverse mean, and moves to a slightly less noisy sample \(x_{t-1}\).
Generation is gradual, not one-shot.
6. Why are Gaussian identities so central?#
Because they give exact formulas for:
the forward marginal \(q(x_t\mid x_0)\)
the posterior \(q(x_{t-1}\mid x_t,x_0)\)
the Gaussian KLs inside the ELBO
Without these identities, the core DDPM derivation would not collapse into such a clean training objective.
Summary#
Core equations in one place#
Forward step#
Forward marginal#
Equivalent sampling form:
Posterior#
with
Reverse model#
Noise-prediction mean parameterization#
Simplified training loss#
Sampling#
Start with
then run the learned reverse chain:
Final conceptual takeaway#
A DDPM is a deep latent-variable generative model built from two coupled processes:
A fixed Gaussian forward process that gradually corrupts data into noise.
A learned reverse process that gradually reconstructs data from noise.
The elegance of DDPMs comes from the fact that Gaussian algebra makes:
the forward marginal tractable,
the posterior tractable,
the ELBO decomposable,
and the practical loss reducible to a denoising regression objective.
That is the mathematical core of DDPMs.
DDPM training from scratch#
Using pure NumPy / Python implementation
# ============================================================
# Minimal DDPM training on pinwheel (pure NumPy / Python)
# ============================================================
# ------------------------
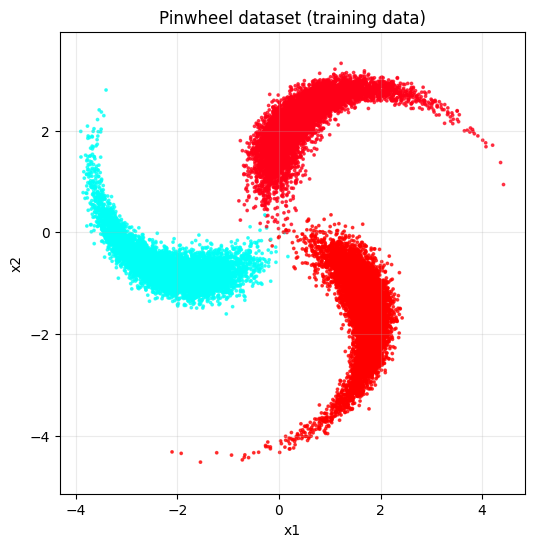
# 1. Create colorful data
# ------------------------
x_np, y_np = make_pinwheel(
n_samples=25000,
radial_std=0.28,
tangential_std=0.10,
num_classes=3,
rate=0.25,
scale=2.3,
)
plot_dataset(x_np, y_np, title="Pinwheel dataset (training data)")
# ------------------------
# 2. DDPM config
# ------------------------
T = 200
schedule = make_ddpm_schedule(T)
model = MLPDenoiser(x_dim=2, time_dim=64, hidden_dim=128, lr=1e-3)
# ------------------------
# 3. Train
# ------------------------
epochs = 25
batch_size = 512
losses = []
for epoch in range(epochs):
epoch_loss = 0.0
n_batches = 0
for x0 in iterate_minibatches(x_np, batch_size=batch_size, shuffle=True, drop_last=True):
bsz = x0.shape[0]
t = np.random.randint(0, T, size=(bsz,), dtype=np.int64)
x_t, noise = q_sample(x0, t, schedule)
loss = model.train_step(x_t, t, noise)
losses.append(loss)
epoch_loss += loss
n_batches += 1
avg_epoch_loss = epoch_loss / max(n_batches, 1)
print(f"Epoch {epoch+1:02d}/{epochs} | avg loss = {avg_epoch_loss:.6f}")
# ------------------------
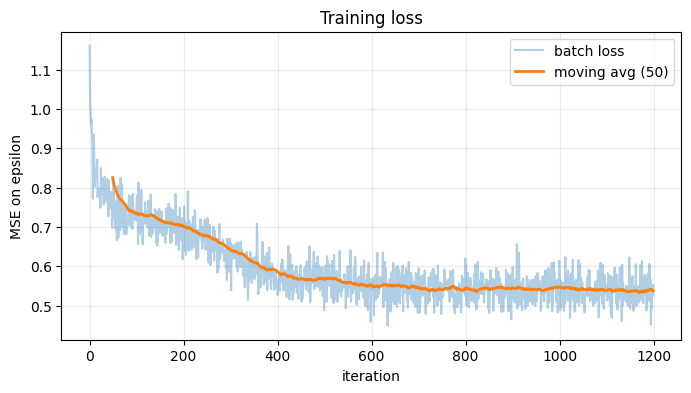
# 4. Plot training loss
# ------------------------
plot_losses(losses, smooth_window=50)
# ------------------------
# 5. Sample from trained model
# ------------------------
samples = sample_final(model, n_samples=6000, schedule=schedule)
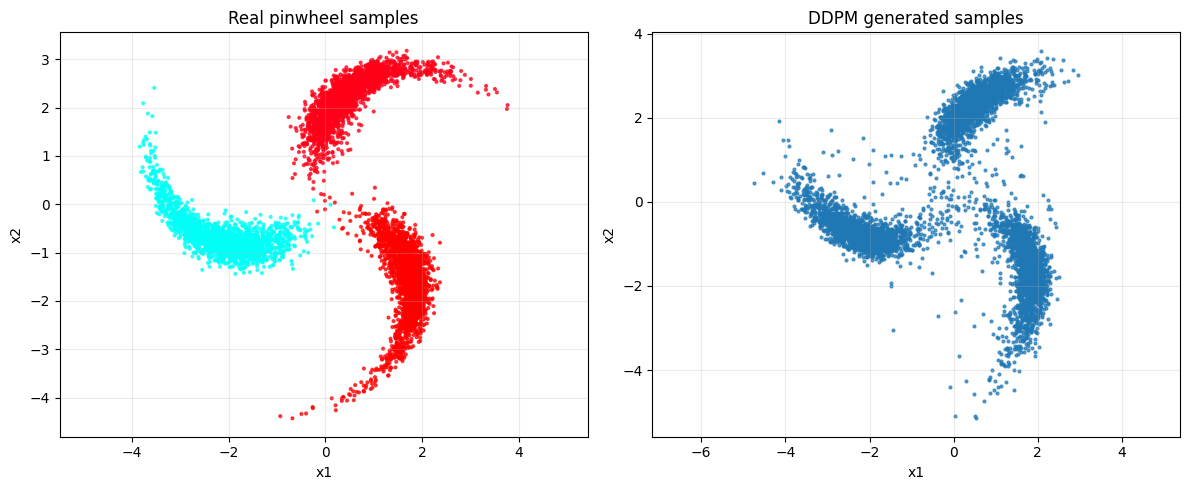
# ------------------------
# 6. Compare data vs generated
# ------------------------
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
axes[0].scatter(x_np[:6000, 0], x_np[:6000, 1], c=y_np[:6000], s=4, alpha=0.7, cmap="hsv")
axes[0].set_title("Real pinwheel samples")
axes[0].set_xlabel("x1")
axes[0].set_ylabel("x2")
axes[0].axis("equal")
axes[0].grid(True, alpha=0.25)
axes[1].scatter(samples[:, 0], samples[:, 1], s=4, alpha=0.7)
axes[1].set_title("DDPM generated samples")
axes[1].set_xlabel("x1")
axes[1].set_ylabel("x2")
axes[1].axis("equal")
axes[1].grid(True, alpha=0.25)
plt.tight_layout()
plt.show()
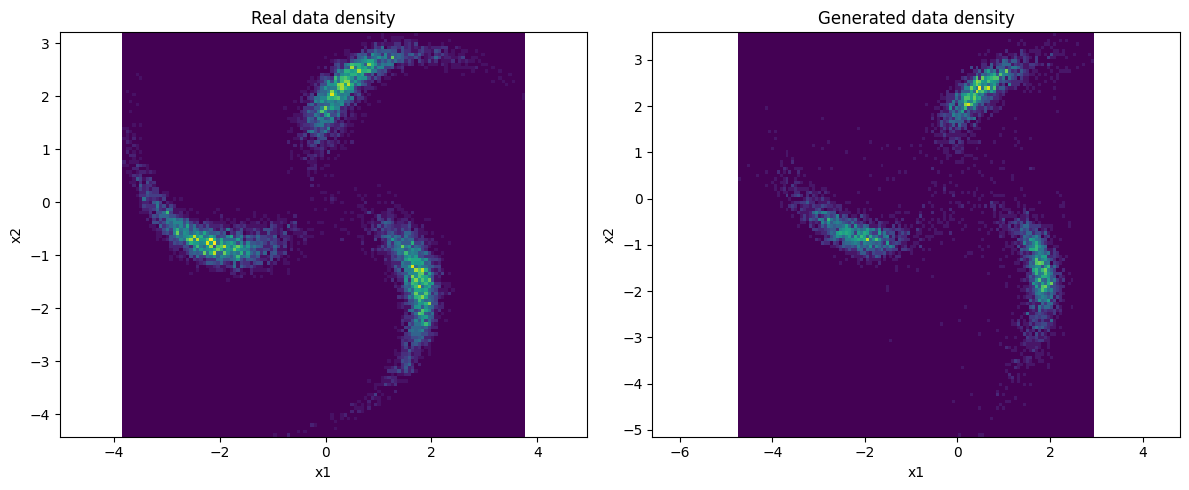
# ------------------------
# 7. Nice density-ish comparison using hist2d
# ------------------------
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
axes[0].hist2d(x_np[:10000, 0], x_np[:10000, 1], bins=120)
axes[0].set_title("Real data density")
axes[0].set_xlabel("x1")
axes[0].set_ylabel("x2")
axes[0].axis("equal")
axes[1].hist2d(samples[:, 0], samples[:, 1], bins=120)
axes[1].set_title("Generated data density")
axes[1].set_xlabel("x1")
axes[1].set_ylabel("x2")
axes[1].axis("equal")
plt.tight_layout()
plt.show()
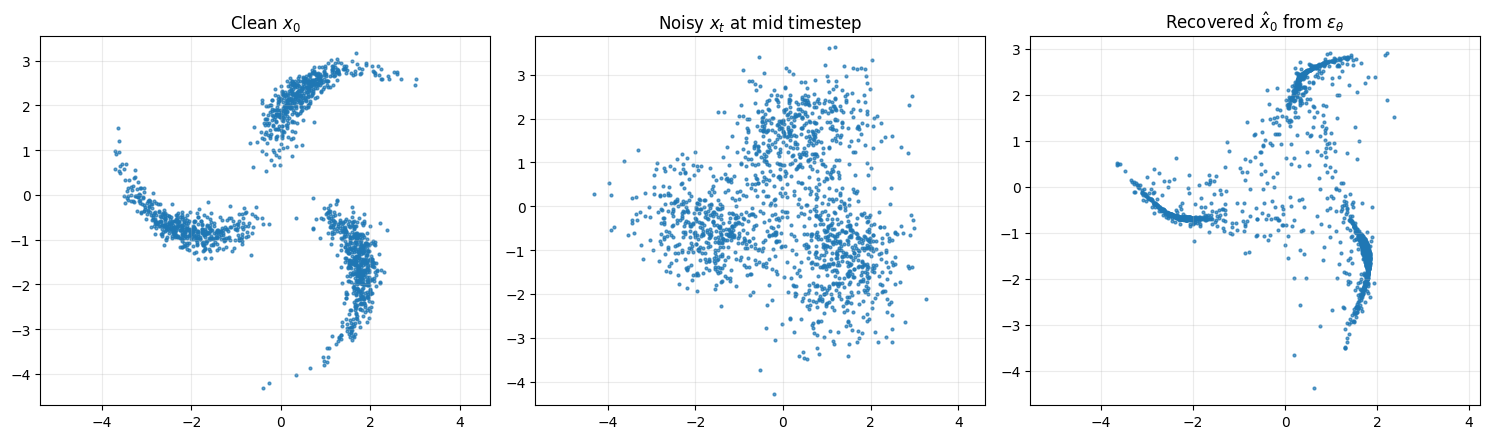
# ------------------------
# 8. Show denoising relation on a random batch
# ------------------------
x0_vis = x_np[:1500]
t_vis = np.full((x0_vis.shape[0],), T // 2, dtype=np.int64)
x_t_vis, eps_true = q_sample(x0_vis, t_vis, schedule)
eps_pred = model(x_t_vis, t_vis)
x0_pred = predict_x0_from_eps(x_t_vis, t_vis, eps_pred, schedule)
fig, axes = plt.subplots(1, 3, figsize=(15, 4.5))
axes[0].scatter(x0_vis[:, 0], x0_vis[:, 1], s=4, alpha=0.7)
axes[0].set_title(r"Clean $x_0$")
axes[0].axis("equal")
axes[0].grid(True, alpha=0.25)
axes[1].scatter(x_t_vis[:, 0], x_t_vis[:, 1], s=4, alpha=0.7)
axes[1].set_title(r"Noisy $x_t$ at mid timestep")
axes[1].axis("equal")
axes[1].grid(True, alpha=0.25)
axes[2].scatter(x0_pred[:, 0], x0_pred[:, 1], s=4, alpha=0.7)
axes[2].set_title(r"Recovered $\hat{x}_0$ from $\epsilon_\theta$")
axes[2].axis("equal")
axes[2].grid(True, alpha=0.25)
plt.tight_layout()
plt.show()
Epoch 01/25 | avg loss = 0.829044
Epoch 02/25 | avg loss = 0.734988
Epoch 03/25 | avg loss = 0.722766
Epoch 04/25 | avg loss = 0.703437
Epoch 05/25 | avg loss = 0.680532
Epoch 06/25 | avg loss = 0.652576
Epoch 07/25 | avg loss = 0.621939
Epoch 08/25 | avg loss = 0.592040
Epoch 09/25 | avg loss = 0.571747
Epoch 10/25 | avg loss = 0.564304
Epoch 11/25 | avg loss = 0.565166
Epoch 12/25 | avg loss = 0.553008
Epoch 13/25 | avg loss = 0.551253
Epoch 14/25 | avg loss = 0.548118
Epoch 15/25 | avg loss = 0.538855
Epoch 16/25 | avg loss = 0.546232
Epoch 17/25 | avg loss = 0.539357
Epoch 18/25 | avg loss = 0.544770
Epoch 19/25 | avg loss = 0.544413
Epoch 20/25 | avg loss = 0.540531
Epoch 21/25 | avg loss = 0.544180
Epoch 22/25 | avg loss = 0.540002
Epoch 23/25 | avg loss = 0.541568
Epoch 24/25 | avg loss = 0.536431
Epoch 25/25 | avg loss = 0.538234